Scientific research is one of the most exciting and rewarding of occupations. It is like a voyage of discovery into unknown lands, seeking not for new territory but for new knowledge. It should appeal to those with a good sense of adventure.
-Frederick Sanger
Scientists are increasingly developing new techniques to investigate the structure and function of DNA. Genes can be taken from organisms and linked to other DNA molecules to form recombinant DNA and introduced into living cells.
A process known as gene cloning, uses the host cell to make many copies of the inserted gene and the protein it codes for. This has allowed the production of transgenic plants and animals; where a gene from one species is introduced into another species. This has brought the possibility of specific gene therapies for human diseases.
Genomics attempts to understand the genome of a species as a whole, including mapping the complete nucleotide sequence for every gene in an organism. This has seen the sequencing of the entire human genome and numerous other mammals, plants, insects and microbes.
MAPPING THE GENOME
Gene mapping involves the idea of fixing a gene to its correct location on one of the chromosomes. This is important to allow further studies of the gene, its inheritance, and possible diagnostic techniques if the gene in question, when mutated, leads to a disease.
In 1990, the U.S. Human Genome Project began and ended in 2003 with a draft version of the sequences of the 3 billion chemical base pairs making up human DNA. This map of the human genome has actually been under construction for the last 100 years beginning with the mapping of the first gene in 1911. This was the gene responsible for red-green colour blindness, which was assigned to the X chromosome following the observation that this disorder was passed on in an X-linked inheritance pattern to sons by unaffected mothers. Subsequently many other disorders affecting only males were likewise mapped to the X chromosome.
The other 22 pairs of chromosomes remained virtually uncharted until the late 1960s, with the discovery of somatic cell hybrids. This involved fusing together human and mouse cells to create unstable cells which quickly lost most of the human chromosomes until only a few remained. Any human proteins in these hybrid cells thus had to be produced by genes located on one of the remaining human chromosomes. This strategy has led to the assignment of around 100 genes to specific chromosomes.
In the 1970’s came the discovery of a staining technique called G-banding, from the Giemsa dye, which made the identification of each chromosome much easier with characteristic darker and lighter bands providing the equivalent of latitudes and serving as rough landmarks on the chromosomes. This led to the assignment of some 1,000 genes to specific chromosomes such as the gene leading to Tay Sachs disease (described in Metabolic Disorders) found on a particular area of chromosome 15.
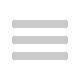
A further development of this technique relied on known variable sequences in the genome sequence as markers for nearby abnormal genes. This was first developed by Kan and Dozy, in 1978, who noted that near the sickle cell gene was a stretch of DNA which varied (i.e. polymorphic) between most Africans and African-Americans and which could be used as a marker in a given family for linkage with the sickle cell gene. This polymorphism served as the first use of these common molecular differences between one person and another as a method for both diagnosis (by being linked to a disease gene) and subsequently for general mapping over the entire genome i.e. finding polymorphic sequences of DNA with no known function (a marker) and linking them with the inheritance of a disease in a family. The identification of these markers was helped by the use of special enzymes found in bacteria, known as restriction endonucleases, which cut DNA at a specific sequence. This technique could be used to analyse very large families to determine which individuals inherited a disease in combination with one of the many different markers on different chromosomes. As such, an explosion in the knowledge of genes’ chromosomal whereabouts occurred with a little less than 2,000 genes being mapped by 1991. This included pinpointing the location of the Huntington’s disease gene (Chromosome 4) using the families in Venezuela, adult polycystic renal disease (Chromosome 16) and cystic fibrosis (Chromosome 7) which were all mapped using this technology.
Meanwhile, scientists learned to sequence these genes from the mid-1970s when Frederick Sanger at Cambridge University and Walter Gilbert and Allan Maxam at Harvard University developed methods for determining the order of bases in a strand of DNA. Automated high-speed sequencing by machine followed in the 1980s, and currently new technology promises to allow the sequencing of an entire genome in a matter of weeks.
DNA FINGERPRINTING
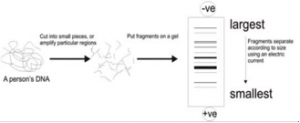
DNA fingerprinting is a technique to allow one to distinguish between individuals using their DNA. Although humans show identical sequences in the vast majority of the genome some specific sequences can be highly variable. It is therefore the likelihood that unrelated individuals would have different sequences at these particular sites in the genome which is the basis for genetic fingerprinting.
An example of such variable sequences are repeats consisting of between 3 and 5 basepairs. When analysing multiple regions containing variable repeats between two individuals a very high statistical power can be generated as one region containing a certain number of repeats does not relate to the number of repeats in any other region. Therefore, if one region in a certain individual has 15 repeats, a second region 30 and a third region 5, the chance of an unrelated person containing the same numbers of repeats is 1 in several million.


This technique can therefore also allow one to determine if two individuals are related. Naturally, identical twins contain identical genetic fingerprints, while relatives will share varying amounts depending on the closeness of the relationship – For this reason, genetic fingerprinting is often used in paternity tests. In 1912 a four-year-old boy called Bobby Dunbar vanished during a fishing trip. Eight months later it was claimed that another boy, Bruce Anderson, living with the family of William Walters, entrusted to them but his mother, was actually the missing child. The courts accepted this story, convicting Walters for the kidnapping and placing the boy with the Dunbar family where he was raised and known as Bobby throughout the rest of his life. However, in 2004 DNA tests on Dunbar’s son and nephew revealed the two were not related, thus establishing that the boy found in 1912 was not Bobby Dunbar, whose real fate remains unknown.
Genetic fingerprinting is also employed in criminal cases to prove that the DNA found at a crime scene is statistically likely to be the same DNA present in a suspect. In 1988, Colin Pitchfork became the first murderer in the UK to be convicted as a result of DNA fingerprinting. Though he had originally evaded police by persuading a friend to give a fraudulent blood sample under his name, he was eventually caught out and sentenced to life in prison for the murder of two young girls. However, controversies do arise from this method, especially when a prosecutor gives some astronomical statistical figure for a DNA sample, which may have been contaminated or mislabelled, belonging to an accused. Indeed, there have been some wrongly ascribed “positive” matches for people who could not possibly have been involved in a crime and it appears several people in the US have been executed on DNA evidence now admitted to be faulty due to human errors in a laboratory. In December 2005 Robert Clark became the 164th person in the USA to be pardoned following post-conviction DNA testing, after serving 24 years of his sentence and in March 2009, Sean Hodgson was released after spending 27 years in jail for murder when tests prove DNA from the scene was not his.
PRENATAL TESTING
There are a number of techniques used to diagnose prenatal defects in foetuses whose mothers are at risk of having a baby with an abnormality.
Amniocentesis involves taking a sample of amniotic fluid (the ‘waters’) using a needle at 15-18 weeks. This can be analysed to determine certain biochemical, chromosomal or neural tube defects such as spina bifida, and can be used to test for specific genetic defects known to be inherited in the family. Another procedure is placental tissue (chorionic villus) sampling, which can be performed earlier at 10 to 12 weeks, but is considered to be more risky to the embryo than amniocentesis.
The most widely used technique is ultrasound scanning using ultrasonic waves to scan the foetus allowing visualisation and enabling skeletal and neural tube defects, among other abnormalities, to be identified. Scans are normally performed between 16 to 20 weeks, but since no actual tissues are sampled the technique cannot include genetic testing.
In the past year groundbreaking research is enabling the possibility to discriminate between tiny samples of foetal DNA in the blood circulation of the mother. This non-invasive technique has been used to diagnose Down Syndrome.
The practice of prenatal diagnosis has recently come under scrutiny, mainly due to different interpretations of what a “serious handicap” is. Many babies in the UK have been, and are being, aborted at advanced pregnancy stages with only mild defects that could easily be corrected, such as polydactyly, club feet and cleft palates. In some countries, notably China, abortion of healthy female babies, though illegal, is not uncommon, leading to a growing gender imbalance in the Chinese population; figures from 2009 show that for every 100 girls born, 119 boys are born.
CLONING
A clone is a single cell (such as bacteria, lymphocytes, etc.) or a multi-cellular organism that is genetically identical to another living organism. Natural clones occur when an organism reproduces asexually or when two genetically identical individuals are produced by accident. The term clone is derived from the Greek word for “twig” which was originally used to describe all descendants of a single plant, produced by vegetative reproduction. In fact many horticultural varieties of plants are clones, with some varieties of grapes representing clones that have been propagated for over 2000 years!
Cloning in the molecular biological sense can refer to two different techniques:
Molecular cloning refers to the procedure where by a piece of DNA is inserted into cells which can then be cultured. This allows the inserted sequence to be analysed, such as testing what protein is derived from any genes which might have been in the region.
Cellular cloning describes the creation of a new organism with the same genetic information as a cell from an existing one. This often refers to the process known as somatic cell nuclear transfer in which a cell of the organism to be cloned, with its nucleus containing the DNA, is transferred into an egg cell which has had its nucleus removed. This results in the host egg cell developing into an organism with genetically identical DNA to the donor.
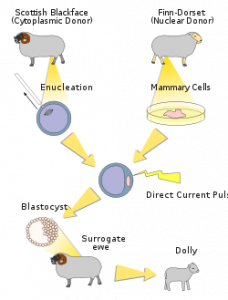

The first animal clone was a frog, cloned by Thomas J. King and Robert W. Briggs in 1952 using somatic cell nuclear transfer. A carp was also created in this way, by inserting DNA of a male carp into the egg of a female carp, in 1963 by an Asian scientist Tong Dizhou, 33 years before Dolly the Sheep. Sadly, like Mendel, he published his findings in an obscure science journal which that was not translated into English. Since then there has been Dolly the sheep (she was cloned from mammary cells hence the name in honour of the country and western singer, Dolly Parton) in 1996 followed by two more sheep, Molly and Polly who were the first transgenic mammals containing a human gene. In 1997 came the arrival of 3 mules by the names of Idaho Gem, Utah Pioneer, Idaho Star (Americans, naturally!), followed by Dewey the deer, Prometea and Paris Texas the horses, 5 Scottish piglets (Millie, Christa, Alexis, Carrel, and Dotcom), an ox named Noah, two Jersey cows named Millie and Emma (there have been several other since), Little Nicky and Copycat the cats, Cumulina the mouse, a rat called Ralph, a rabbit which was sadly nameless (developed by the French), a dog named Snuppy, a monkey called Tetra and, supposedly, a human called Eva.
It was in December 2002 that Clonaid, the medical arm of a cult called Raëlism who believe that aliens introduced human life on Earth, claimed to have successfully cloned the first human being, a baby named Evá. They maintained that aliens taught them how to perform cloning, even though the company had no record of having successfully cloned any previous animal. The company has failed to provide the DNA which would allow an independent agency to prove that the girl was a clone and their claims are now generally discredited.
In 2008 five human embryos were created using DNA from skin cells, aiming to provide a source of viable embryonic stem cells. However, the embryos were destroyed at a very early stage of development.
STEM CELLS
Stem cells are undifferentiated cells that retain the ability to divide and differentiate into many other cell types. These cells are found naturally in certain areas of the adult body, and in the embryo, where they serve to replenish cells in the body. The main types of stem cells are the embryonic stem cells and adult stem cells which differ in their “potency”. Adult stem cells are described as “multipotent” meaning that they can develop into closely related cells though are unable to differentiate into every cell type in the human body. For example, hematopoietic stem cells can only differentiate into red blood cells, white blood cells and platelets. Embryonic stem cells, on the other hand, are described as being pluripotent in that they can form any of the 200 cells of the adult human body.
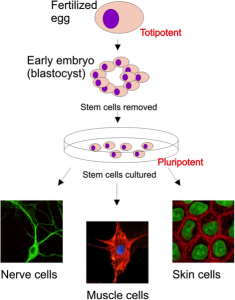
Embryonic stem cells are cells derived from the inner cell mass of developing blastocysts. An ES cell is self renewing (can replicate itself), pluripotent (can form all cell types found in the body) and theoretically is immortal.
These embryonic stem cells are found only in the very early stage embryo which, in humans, are approximately 4 to 5 days old and consist of 50 to 150 cells. Because of their unique combined abilities of unlimited expansion and pluripotency, embryonic stem cells are a potential resource for regenerative medicine and tissue replacement after injury or disease. Medical researchers anticipate being able to use technologies derived from stem cell research to treat cancer, Parkinson’s disease, spinal cord injuries, and muscle damage, amongst a number of other diseases, impairments and conditions. However, there is widespread controversy over stem cell research relating mainly to the way in which stem cells are acquired from human embryos resulting in the destruction of a possible human life. Although President George W. Bush became the first president to provide federal funding for embryonic stem cell research, totalling approximately $100 Million, in July 2006 he vetoed a bill that would have allowed this money to be used for studies involving the destruction of embryos. New research, however, shows that it is possible to alter certain genes within a normal non-stem cell allowing a differentiated cell to regain its pluripotency.
GENE THERAPY
Gene therapy is the insertion of genes into an individual’s cells and tissues to treat genetic diseases by supplementing a defective mutant allele with a functional one.
In the 1980s, methods became available to allow easy ways to produce proteins, such as the insulin protein, deficient in diabetics, or clotting factors missing in patients with haemophilia. This is done by introducing human genes into bacterial DNA, so that the modified bacteria can then produce the corresponding protein, known as a recombinant protein, which can be harvested from the bacteria and injected in people who cannot produce it naturally. Scientists took the logical step of trying to introduce genes straight into human cells, focusing on diseases caused by single-gene defects, such as cystic fibrosis, haemophilia, muscular dystrophy and sickle cell anaemia.
Target cells such as the patient’s liver or lung cells are infected with a viral vector, used to carry genetic material containing the therapeutic human gene, into the target cell. The generation of a functional protein product from the therapeutic gene restores the target cell to a normal state.
In theory it is possible to transform either somatic cells (most cells of the body) or cells of the germline (such as sperm cells, ova, and their stem cell precursors). All gene therapy so far in people has been directed at somatic cells, whereas germline engineering in humans remains only a highly controversial prospect since the introduced gene could be transmitted to offspring. Somatic gene therapy can be broadly split in to two categories: ex vivo (where cells are modified outside the body and then transplanted back in again) and in vivo (where genes are changed in cells still in the body).
On September 14, 1990 researchers in the U.S. performed the first approved gene therapy procedure on four-year old Ashanti DeSilva who was born with SCID. The procedure involved removing white blood cells from her body, inserting the missing gene into the cells, and then infusing them back into her bloodstream. Tests showed that the therapy strengthened Ashanti’s immune system though she needed to continually receive replacement gene therapy. Therefore, while this procedure is not a cure it did prove that in theory the symptoms could be alleviated. However trials in France highlighted the possible pitfalls to such a procedure with a number of children treated for XSCID with gene therapy, developing leukaemia due to the randomly inserted gene sometime disrupting further genes leading to cancer. As a result of this, gene therapy treatments for SCID were discontinued for a long period of time. However newer approaches combined with using different modified viruses to introduce genes into cells are beginning to reopen the door to treating a range of diseases from haemophilia, adrenoleukodystrophy and some forms of blindness.
